Free Software Coral
Coral(CORrelation And Logic).
The archive (CORALSEA.zip) contains folders:
(i) examples of databases;
(ii) the software for regression models and for classification models;
(iii) CORALSEA: An updated version that combines the features of the previous ones, as well as some new ideas.
You can download the zip-file from here.
There is a collection of models built with CORAL software.
To view this collection as well as calculate endpoint values using SMILES you are interested in,
click "MODELS".
The detailed description of the CORALSEA
software you will find in «ReadMe file» as well as in
«Brief illustrations and instructions»,
where difference between regression model and classification model is clarified.
The archive (CORALSEA.zip)
contains databases on physicochemical parameters (e.g.octanol/water
partition coefficient, enthalpy of
formation from elements, solubility) and biological activity (e.g.
toxicity, mutagenicity, carcinogenicity).
These data are available for
new computational experiments with CORALSEA.
In principle, these data can be involved in computational experiments with
other software, which can use SMILES as the representation of the molecular structure.
One can use these models for prediction of endpoint values for external substances (these should be similar to substances involved in the models) as well as these can be used as templates to build
up new models using other data in the same format.
 New options:
New options:
SOFTWARE WITH ATTRACTIVE ADDITIONAL POSSIBILITIES!
CORALSEA-2016
1. Automatic distribution into training and validation sets
2. Detection of duplicates
3. New Super-attribute of SMILES
4. Design of mechanistic interpretation
5. Control for input files
CORALSEA-2017(1)
IIC, CCC and CII (CORAL-2020):
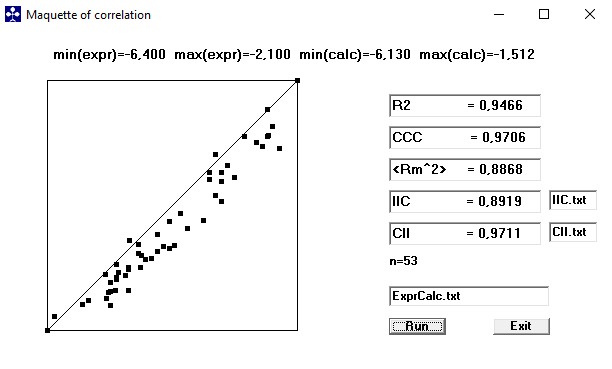
The Index of Ideality of Correlation (IIC), the Concordance Correlation Coefficient (CCC) and the Correlation Intensity Index (CII-2020).
These new criteria of predictive potential (IIC, CCC and CII) are available in autonomic program. Test these with any models (not only CORAL models) with using our program! You can download the IIC.zip-file from here.
CORALSEA-2017(2)
IDENTITY OF SPLITS:
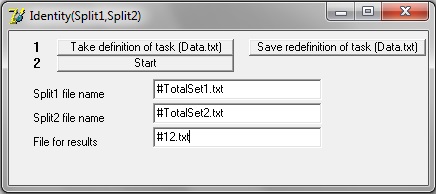
In order to estimate, identity of splits (into training set ... validation set) you can use special autonomic program which can be downloaded
here.
CORALSEA-2019
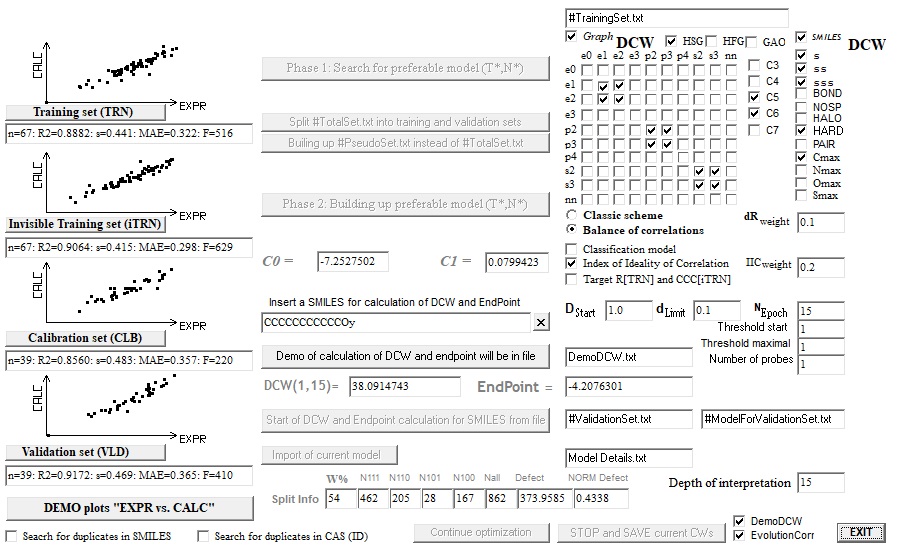

The difference of CORALSEA-2019 in comparison with CORALSEA-2017 contains the following:
1. The "Square of binary combinations of topological invariants" of molecular graph
2. Additional SMILES attributes: Cmax, Nmax, Omax, and Smax
3. Possibility to stop the Monte Carlo optimization
4. Possibility to continue the Monte Carlo optimization
An example of application of CORALSEA-2019 is available:
J. Mol. Struct., 1182 (2019) 141-149.
https://doi.org/10.1016/j.molstruc.2019.01.040
CORALSEA-2020
The difference of CORALSEA-2020 in comparison with CORALSEA-2019 contains the following:
1. The Correlation Intensity Index (CII) is a new criterion of predictive potential. In CORALSEA-2020, the CII can be involved in the Monte Carlo optimization. Preliminary computational experiments confirm that the CII improves statistical quality of some models.
2. Self-organized atoms pairs proportions vector. Atoms pairs proportions is extended version of atoms pairs available in CORALSEA-2016 and CORALSEA-2017. These SMILES attributes reflects not only presence of corresponding atoms, but also their rations (1:1, 1:5, 3:7, etc.).
3. Possibility to apply new types of the Monte Carlo optimization.
4. New options for design of the quasi-SMILES codes.
5. The calculation of the values of the endpoint using the current model for a list of the "dark SMILES" (or "dark quasi-SMILES").
6. Graph invariants indicated in CORAL screen by red. SMILES attributes indicated by green. This can be convenient for potential users.
7. Updated software provides users by the statistical status of SMILES (falls or not in domain of applicability) for all sets (active training, passive training, calibration, and validation sets).
CORALSEA-2023 New possibilities: new idealisation, new simplicity!
CORALSEA-2025 New: Las Vegas algorithm!
During 2024-2025, a procedure was developed to find "convenient" splits into training and validation sets. It is assumed that the training set is structured into active and passive training sets, supplemented by a calibration set. The above procedure is based on the Las Vegas algorithm, which produces the preferable split for the calibration set from a group of runs of Monte Carlo optimization. It is assumed this set should also be good for the external validation set (instructions included).